Spatial Intelligence for Biological Systems
Develop multimodal AI methods to model cellular interactions, tissue architecture, and spatial ecosystems across imaging and omics modalities.
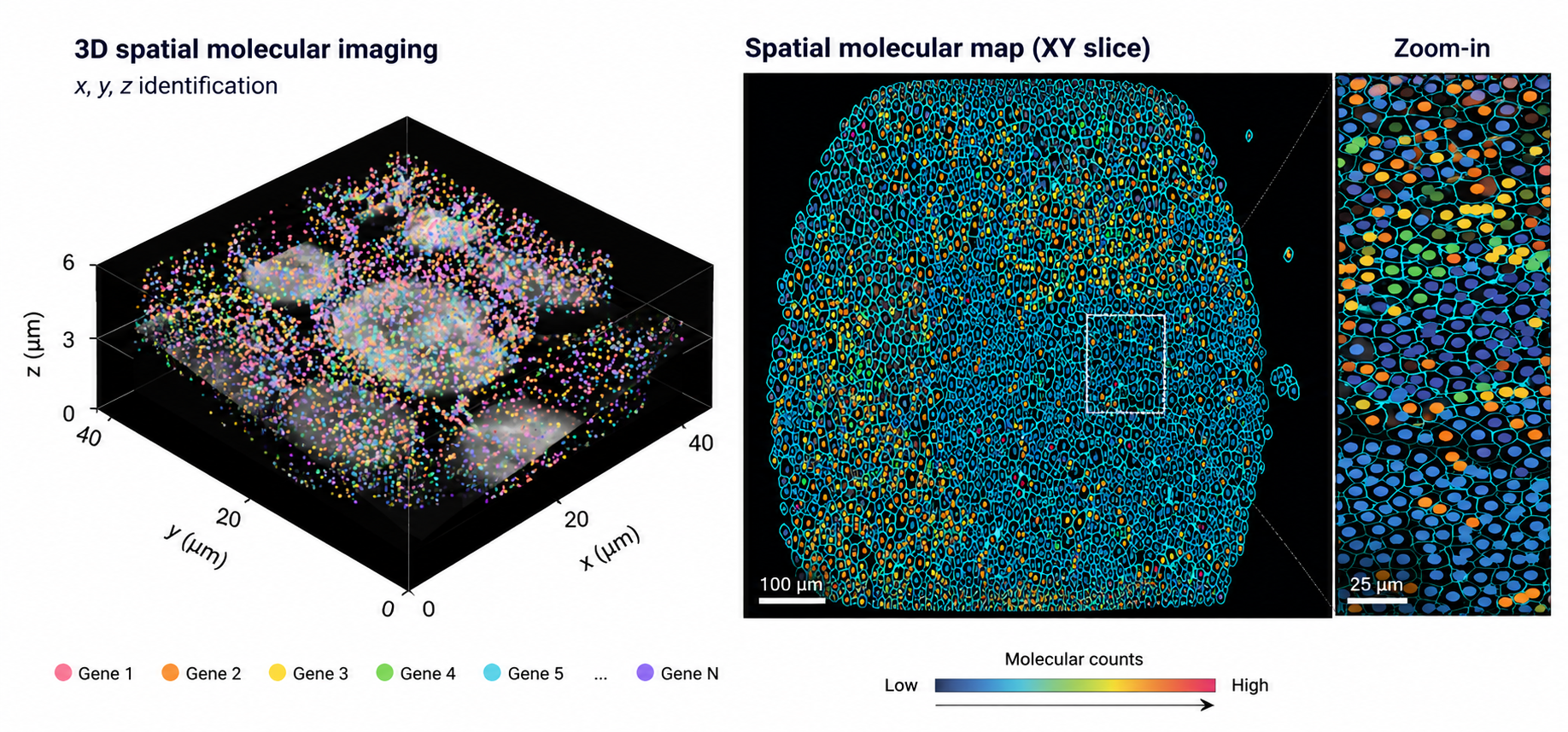

Spatial Intelligence · Computational Medicine · Imaging and Signal processing
Spatial AI Intelligence
We develop next generation multi-modal AI for modeling spatially organized biological and physical systems, integrating imaging, omics, and sensing data to advance healthcare, energy, and environmental sciences.
Research
Develop multimodal AI methods to model cellular interactions, tissue architecture, and spatial ecosystems across imaging and omics modalities.

Build large-scale representation learning and foundation models for medical imaging, spatial omics, and clinical language data.
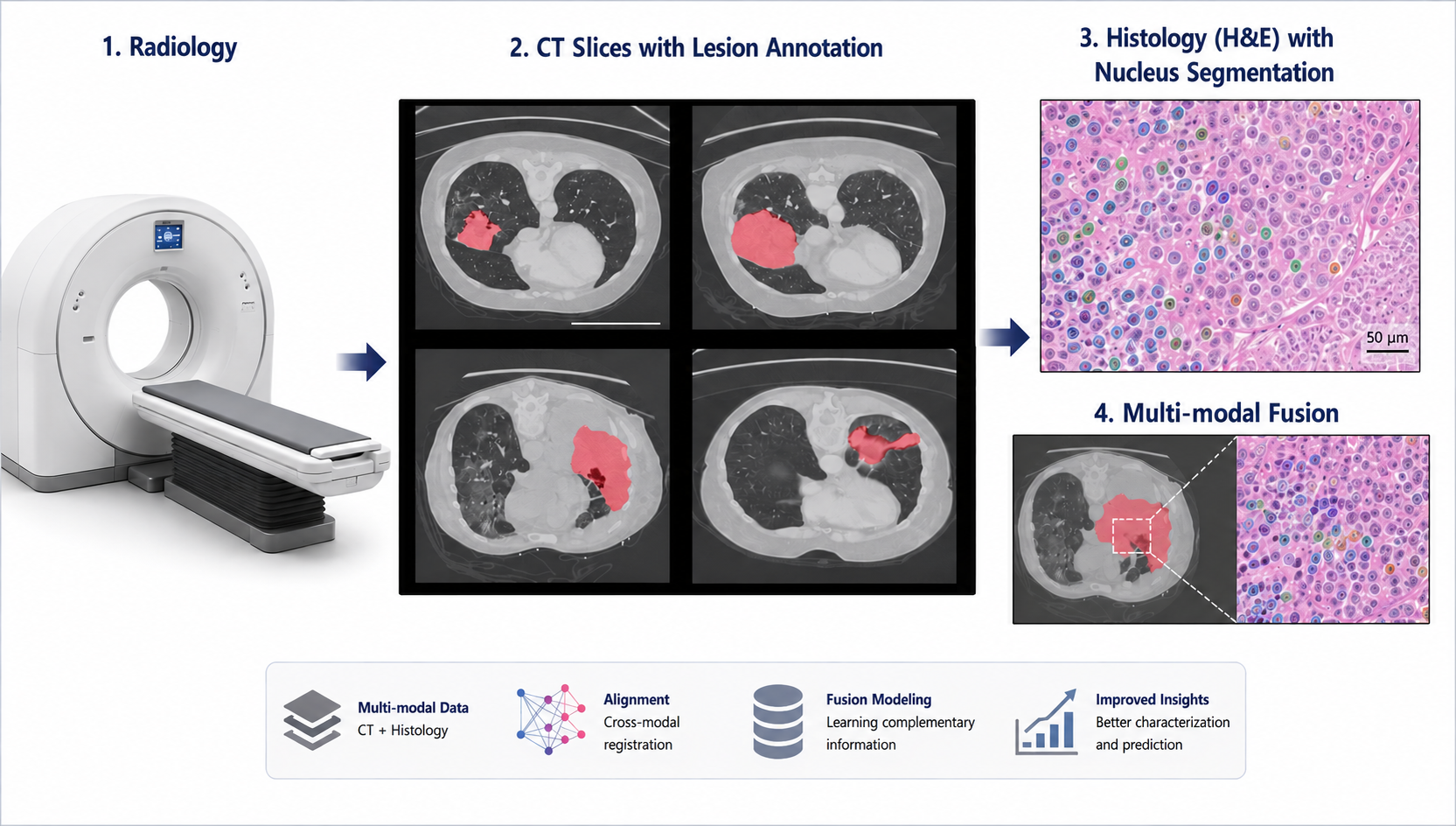
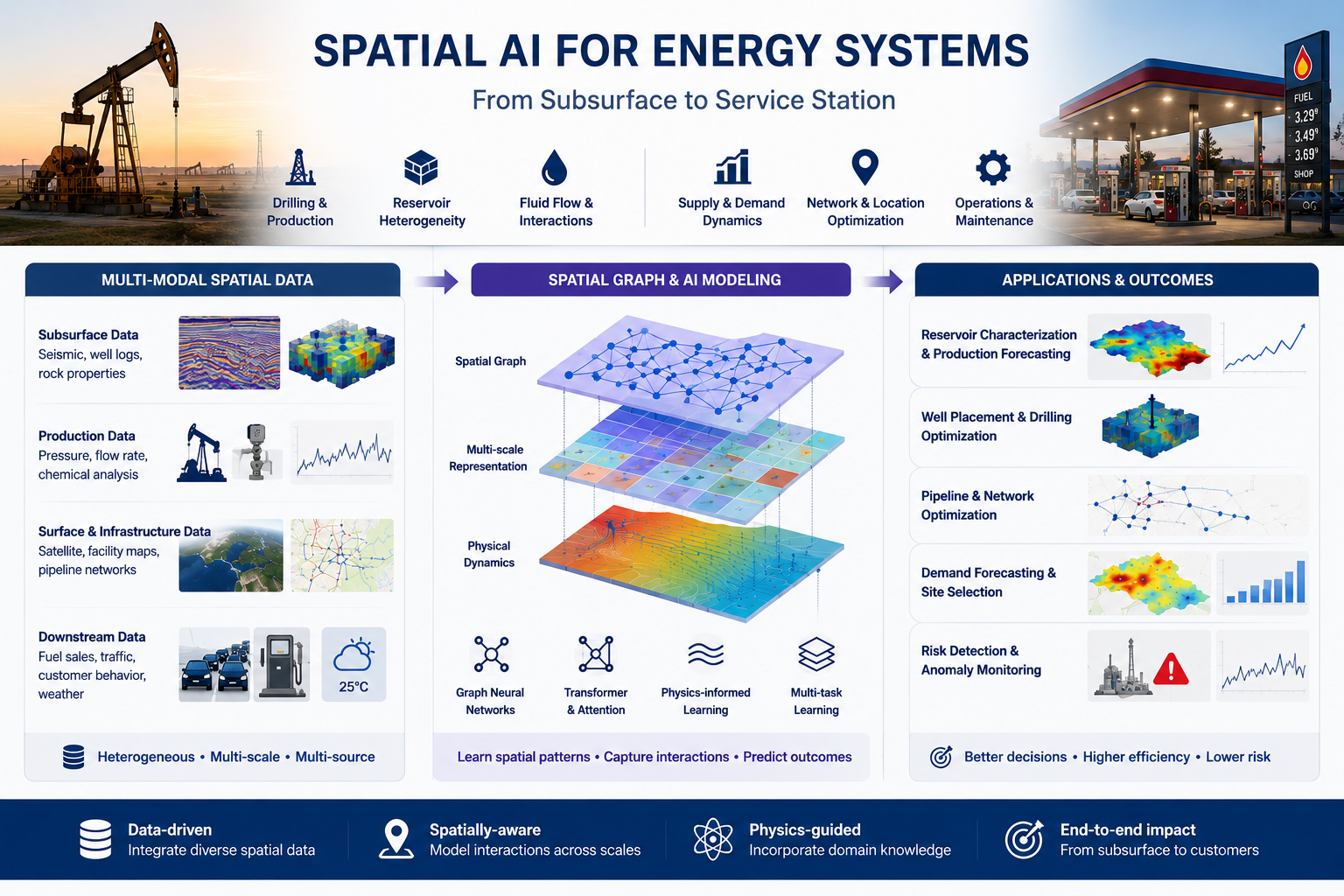
Develop AI frameworks for multimodal sensing, representation learning, and optimization in complex physical and energy systems.
Principal Investigator
Dr. Yuanning Zheng is a translational AI scientist whose research focuses on developing next-generation intelligent systems to advance precision medicine, healthcare, and sustainable energy systems. His work integrates multi-modal artificial intelligence, spatial modeling, medical imaging, and large-scale biological and physical data to better understand complex systems across healthcare and engineering domains.
Dr. Zheng received his M.S. in Computer Science from Georgia Institute of Technology and his Ph.D. from Texas A&M University. He completed his postdoctoral training at Stanford University, where he was mentored by Olivier Gevaert and Zinaida Good.
His research has been supported by the NIH/NCI K99/R00 Pathway to Independence Award. Dr. Zheng maintains active collaborations with clinicians, pathologists, engineers, and AI scientists across academia and medicine, with the goal of translating computational innovations into real-world impact in healthcare, energy, and environmental systems.
Contact: [email protected] LinkedIn Google Scholar ORCID
Software
Selected publications
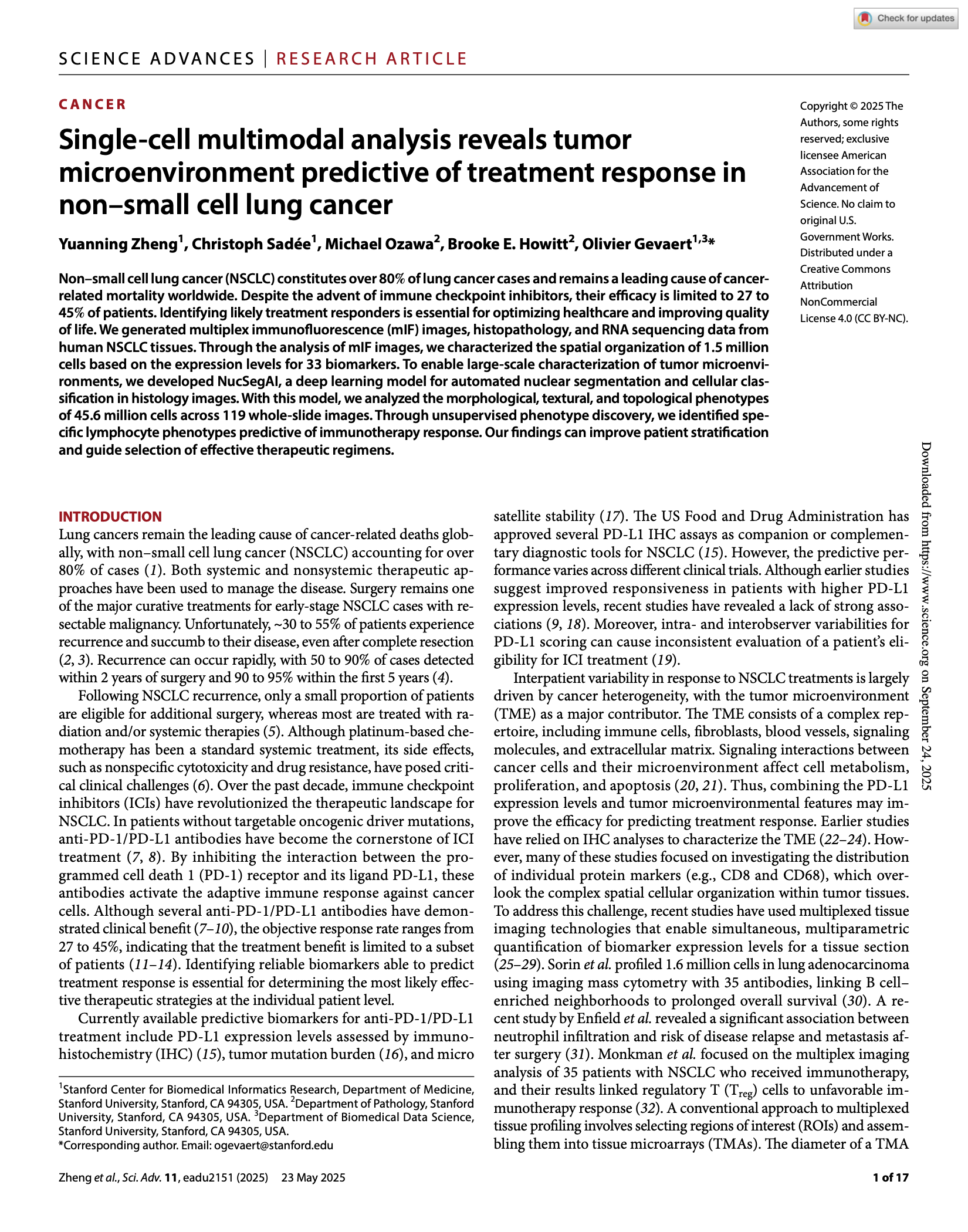

Zheng Y, Sadée C, Ozawa M et al. Single-cell multi-modal analysis reveals tumor microenvironment predictive of treatment response in non-small cell lung cancer. Science Advances, 11(21), 2025.

Pizurica M*, Zheng Y*, Carrillo-Perez F et al. Digital profiling of cancer transcriptomes from histology images with linearized vision attention. Nature Communications, 15(9886), 2024.

Brooks J*, Zheng Y*, Hunter K et al. Digital Spatial Profiling identifies distinct patterns of immuno-oncology-related gene expression within oropharyngeal tumours in relation to HPV and p16 status. Frontiers in Oncology, 14, 2024.

Zheng Y, Carrillo-Perez F, Pizurica M et al. Spatial cellular architecture predicts prognosis in glioblastoma. Nature Communications, 14(4122), 2023.

Zheng Y, Jun J, Brennan K et al. EpiMix is an integrative tool for epigenomic subtyping using DNA methylation. Cell Reports Methods, 3-100515, 2023.

Thieme AH, Zheng Y, Machiraju G et al. A deep-learning algorithm to classify skin lesions from mpox virus infection. Nature Medicine, 29, 738-747, 2023.

Carrillo-Perez F, Pizurica M, Zheng Y et al. Generation of synthetic whole-slide image tiles of tumours from RNA-sequencing data via cascaded diffusion models. Nature Biomedical Engineering, 2024.

Nourbakhsh M, Zheng Y, Noor H et al. Revealing cancer driver genes through integrative transcriptomic and epigenomic analyses with Moonlight. PLOS Computational Biology, 21(4), e1012999, 2025.
Zhan X, Xu Q, Zheng Y et al. Reliability-based cleaning of noisy training labels with inductive conformal prediction in multi-modal biomedical data mining. PLOS Computational Biology, 21(2), e1012803, 2025.

Noor H, Zheng Y, Mantz A et al. A 20-feature radiomic signature of triple-negative breast cancer identifies patients at high risk of death. npj Breast Cancer, 11-79, 2025.

Steyaert S, Qiu YL, Zheng Y et al. Multimodal deep learning to predict prognosis in adult and pediatric brain tumors. Communications Medicine, 3-44, 2023.

Bareja R, Carrillo-Perez F, Zheng Y et al. Evaluating Vision and Pathology Foundation Models for Computational Pathology: A Comprehensive Benchmark Study. Nature Communications, accepted.
Team
We are recruiting highly motivated PhD students, postdoctoral fellows, and research staff who are passionate about developing next-generation spatial intelligence and multi-modal AI to advance precision medicine, energy systems, and environmental science. Successful candidates will have:
Candidates with backgrounds in artificial intelligence, machine learning, computational biology, computer vision, medical imaging, bioinformatics, data science, electrical engineering, or related quantitative disciplines are highly encouraged to apply.
Interested applicants should send the following materials to the PI's email address:
Principal investigator
Spatial AI, multimodal biomedicine, digital pathology, and translational AI.
[email protected]PhD student
Future trainee in spatial AI for complex biological and physical systems.
Postdoctoral scholar
Future researcher in foundation models, imaging processing, and translational AI.
Software engineer
Future builder of robust research software, model infrastructure, and data platforms.
Alumni
Contact
We welcome collaborations and discussions with clinicians, pathologists, biologists, engineers, and industrial partners on multimodal AI, spatial intelligence, medical imaging, complex biological and physical systems, and translational applications in healthcare, energy, and environmental science.